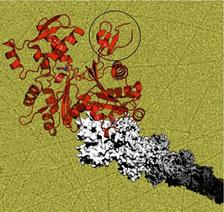
Structure of the plasmodium protein actin I (red). Replacing the D-loop in subdomain 2 (circled) with that of vertebrate muscle actin causes actin I to form long filaments (grey and background) instead of the usual short oligomers. (Credit: Inari Kursula/CSSB)
An international team of scientists has decoded two key proteins of the malaria parasite Plasmodium, by using PETRA III and other facilities. The results shed light on the workings of Plasmodium's structural proteins actin I and actin II, without which the parasite cannot infect human cells. The project led by Prof. Inari Kursula from the new Centre for Structural Systems Biology (CSSB) on the DESY campus may contribute to the development of tailor-made drugs against malaria. The researchers present their work in the open access scientific journal "PLoS Pathogens".
25 April 2014 has been declared World Malaria Day by the World Health Organization (WHO). Malaria is a life-threatening disease that causes more than half a million deaths per year. According to WHO estimates, there were about 207 million cases of malaria in 2012. Children in Africa are at an especially high risk, and there is no approved vaccination to date. The disease is caused by Plasmodium parasites–single-celled parasites, which are transmitted by mosquitos. The pathogen enters the human body through a bite and induces typical symptoms like periodic fevers, nausea, and headaches.
To enter human cells and leave them again, the parasites need to be motile. To this end, they use a structural protein called actin. Actin is found in nearly all living organisms where it is one of the most abundant proteins. Inside cells, it assumes numerous tasks: It confers stability, allows cell division, and makes movement of single cells possible. The dynamical behaviour needed for these processes is enabled by individual globular actin molecules assembling together to form thread-like structures called filaments. The malaria parasite possesses two versions of actin, actin I and actin II, which differ substantially from each other. Even though these structural proteins are crucial for the pathogen’s infectivity, researchers have so far not been able to demonstrate filament formation in the parasite.
Scientists of the Helmholtz Centre for Infection Research (HZI), the European Molecular Biology Laboratory (EMBL) and DESY - three of the nine partner institutions in the CSSB - together with international partners, now succeeded in detecting filament assembly of the parasite actin II proteins. For this, they used electron microscopy, which overcomes the resolution limit of classical light microscopy. Male malaria parasites from which the scientists had deleted actin II were not able to form mature germ cells and consequently could not reproduce and propagate. To have only one actin variant is apparently not sufficient for this process. How filaments contribute to germ cell maturation is still unclear. But why do the two proteins show such different behaviour?
To answer this question, the research team deciphered the structure of the globular actin proteins using bright X-rays at the PETRA III experimental station P11 and at other facilities. “We were able to determine the structures of actin I and actin II at very high resolutions–down to 1.3 and 2.2 Ångström, respectively. With this, we are in the range of single atoms,” says Kursula. “The structures show us that the two variants differ more from each other than actins in any other known living organism do.”
The high resolution at PETRA enabled the researchers to identify areas within the proteins that cause the different behaviour. “We now understand that Plasmodium actin filaments are very different from other actin filaments, like for example from those found in humans, and that they are assembled in a very different manner. Now that we know the structural basis for this, we can look for ways to specifically interfere with the parasite cytoskeleton,” says Kursula. This knowledge might in the future contribute to designing tailor-made anti-malarial medication.
CSSB on the DESY campus is a cooperation of nine research institutions, including DESY, HZI and EMBL.
(from DESY News / HZI press release)
Original publication
"Structural Differences Explain Diverse Functions of Plasmodium Actins"; Juha Vahokoski et al.; PLOS Pathogens, 2014; DOI: 10.1371/journal.ppat.1004091
World Malaria Day: http://www.who.int/campaigns/malaria-day/2014/en/