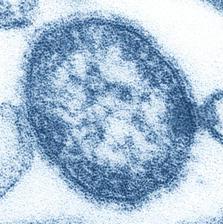
Thin-section transmission electron micrograph (TEM) of a single virus particle, or “virion”, of measles virus. The measles virus is a paramyxovirus. It is 100-200 nanometres in diameter. (Credit: CDC)
Researchers from Ludwig-Maximilians-University and the Center for Integrated Protein Sciences in Munich, Germany, have uncovered how paramyxoviruses, such as the measles virus, attempt to evade an organism’s inborn immune system. Receptor molecules, an important part of the immune system, are able to sense a broad range of viruses. Paramyxoviruses, in turn, produce what is known as V protein, which binds to the immune receptor MDA5 and limit the cell’s immune response. Their work provides detailed insights into the mechanisms how viral proteins can inhibit host protein functions. It may also open up opportunities for new therapeutic interventions.
The research team obtained key structural information on V protein-binding to MDA5 through X-ray experiments performed at the ESRF (Grenoble, France) and at DESY’s lightsource DORIS. The scientists reported their findings in the journal Science.
The antiviral immune system protects an organism against diseases by recognizing viral molecules and initiating an immune response. The immune receptor protein MDA5, a member of the family known as “RIG-I like receptors”, recognizes viral RNA – the intruder’s genetic material – by assembling into filaments along the RNA. It is believed that this filament formation triggers antiviral signaling inside the cell. However, viruses have evolved survival strategies, enabling them to play “hide-and-seek” with the immune system. The virus that causes measles, for instance, produces a so-called V protein, which binds specifically to the RIG-I like receptors MDA5 and LGP2 and thus impairs recognition of virus-infected cells by the adaptive immune system. However, it does not inhibit RIG-I itself. Fortunately, additional cellular recognition mechanisms are in place, limiting the virus’ virulence.
To study MDA5 inhibition in detail, structural information on the interactions between the V and MDA5 proteins is essential. They were able to crystallize the complex formed by the V protein and MDA5 for the first time and determine its three-dimensional structure. With X-ray crystallographic data from the ESRF, the researchers established that the V protein partially unfolds MDA5 via a hairpin-like structure and binds to a site normally buried inside the host protein. Unfolding of MDA5 disrupts filament formation along the RNA and hence antiviral signaling.
In the crystallographic analysis, both proteins were truncated in length, and the crystal structure focused on protein constructs large enough to study protein-protein interactions. The researchers state that their structure “represents the evolutionary conserved, necessary and sufficient core of the MDA5:V complex. Nevertheless, protein domains not present in the crystal structure are likely to impact the overall flexibility of the entire complex, i.e. formed by non-truncated proteins. From a structural comparison with a different V protein-involving complex and from mutational studies of the full-length V protein, the researchers concluded that not only does MDA5 undergo a conformational change in the formation of the MDA5:V complex, but also the V protein itself. In the absence of MDA5, the V protein’s hairpin structure is bound to another V protein domain. In the presence of MDA5, however, the hairpin unfolds and binds to MDA5, which then also unfolds.
To further support their model, the researchers conducted small-angle X-ray scattering (SAXS) experiments at the DORIS beamline X33 , operated by the European Molecular Biology Laboratory (EMBL). Whereas crystallography yields high-resolution structural models of protein crystals, the SAXS technique enables scientists to determine low-resolution protein shapes and protein interactions in solutions. They routinely used SAXS to complement our crystallographic studies. Their crystal structure showed only part of the MDA5:V complex. However, the SAXS studies on the full-length proteins validated the model and we obtained insights into the shape and flexibility of the entire complex.
In their study, the scientists also explained why the V protein specifically binds to MDA5 (and LGP2) but not to the related antiviral RNA sensor RIG-I. The very small hairpin structure of the V protein interacts with MDA5 by making two characteristic sequence-specific contacts. In other words, most contacts between the hairpin and MDA5 do not depend on the amino acid sequences of the protein backbones except for two unique amino acid pairs. Exchanging only two amino acids in RIG-I, which normally does not bind the V protein, allows us to engineer robust binding.
The research may also have possible biomedical implications. With the structural information available, scientists could attempt inhibiting the interactions between the V and MDA5 proteins, thereby reducing the virus’ virulence.
(from DESY News)
Paramyxovirus V Proteins Disrupt the Fold of the RNA Sensor MDA5 to Inhibit Antiviral Signaling; Carina Motz, Kerstin Monika Schuhmann, Axel Kirchhofer, Manuela Moldt, Gregor Witte, Karl-Klaus Conzelmann, Karl-Peter Hopfner; Science, 2013,